One of the major contributors to improved healthcare outcomes and reduced mortality in the post-World War II era was the widespread use of antibiotics. Able to treat infection, these miracle drugs literally transformed bacteria from humanity’s primary killer to a manageable problem.
But, in recent decades, the decline in the discovery of novel antibacterials (and other antimicrobials like antifungals) and the rapid rise in antimicrobial resistance has threatened this happy state. This has led many experts to worry about what will happen in a post-antibiotic world. After all, without effective antibiotics, we not only lose the ability to treat life-threatening diseases like tuberculosis (not to mention common ailments which are a nuisance today but may become more serious like strep throat, urinary tract infections, and ear infections), but we will also lose the ability to perform many surgeries safely (as recovery oftentimes counts on antibiotics to prevent and hold at bay infections).
While we need to get smarter about how and where we use antibiotics (especially in agricultural applications) to slow the rise of resistance, the other half of this problem is in discovering and commercializing new antimicrobials. This is something we’ve largely failed to do since the 1960s, as the figure below from the C&EN article derived from data in a 2019 paper shows.
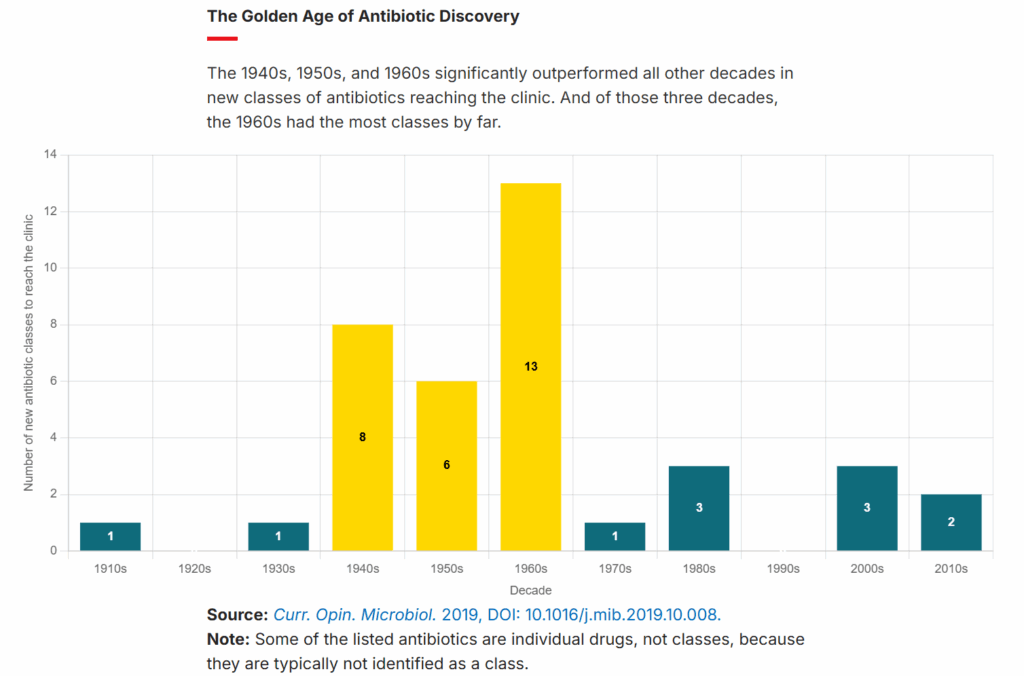
The “golden age” of antimicrobial discovery that ended in the 1960s came largely from researchers searching for these miracle chemicals in soil samples (“bioprospecting”), where bacteria and fungi, in order to compete, release them as “chemical weapons” against others. But, having long ago exhausted the “easy” antimicrobials, we were unable to replicate this success in the decades following the 1960s
But maybe this is about to change. In March, two completely new classes of antimicrobial were published in two weeks (both in Nature) — equaling the total output of the 2010s! Both papers (one by a group from China Pharmaceutical University in Nanjing, China and the other from a collaboration between University of Illinois in Chicago and McMaster University in Canada) were centered around identifying biosynthetic gene clusters (BGCs) responsible, and with the lower cost of metagenomic sequencing and new computational methods, this may point to a new era of discovering novel antimicrobials to help us innovate our way out of our current antimicrobial resistance dilemma by re-inventing bioprospecting.
A good reminder that what we need is the political and scientific willpower to keep funding this type of research and the types of genomic and protein databases that make this type of innovation possible!
In March, the journal Nature published the discoveries of two paradigm-breaking antimicrobial compounds in as many weeks: a polyene macrolide antifungal called mandimycin and a lasso peptide antibiotic called lariocidin.
Both these compounds, which use never-before-seen antimicrobial mechanisms, were found using techniques that let researchers look deep into the chemical diversity of microbes—much deeper than a typical antibiotic or antifungal screen might go. And it’s not just these two molecules. Scientists are using the new approaches to discover countless other antimicrobial compounds with the potential to become drugs.

Bioprospectors mine microbial genomes for antibiotic gold
Max Barnhart | C&EN