The paper I read is something that is very near and dear to my heart. As is commonly known, individuals of Asian ancestry are more likely to experience dizziness and flushed skin after drinking alcohol. This is due to the prevalence of a genetic defect in the Asian population which affects an enzyme called Aldehyde Dehydrogenase 2 (ALDH2). ALDH2 processes one of the by-products of alcohol consumption (acetaldehyde).
In people with the genetic defect, ALDH2 works very poorly. So, people with the ALDH2 defect build up higher levels of acetaldehyde which leads them to get drunker (and thus hung-over/sick/etc) quicker. This is a problem for someone like me, who needs to drink a (comically) large amount of water to be able to properly process wine/beer/liquor. Interestingly, the anti-drinking drug Disulfiram (sold as “Antabuse” and “Antabus”) helps alcoholics keep off of alcohol by basically shutting down a person’s ALDH2, effectively giving them “Asian alcohol-induced flushing syndrome” and making them get drunk and sick very quickly.
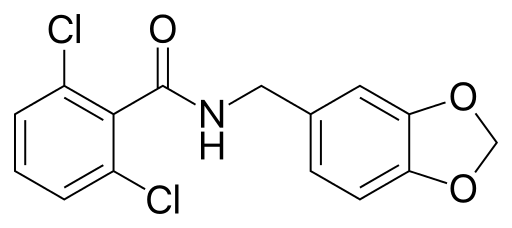
So, what can you do? At this point, nothing really (except, either avoid alcohol or drink a ton of water when you do drink). But, I look forward to the day when there may actually be a solution. A group at Stanford recently identified a small molecule, Alda-1 (chemical structure above), which not only increases the effectiveness of normal ALDH2, but can help “rescue” defective ALDH2!
Have we found the molecule which I have been searching for ever since I started drinking? Jury’s still out, but the same group at Stanford partnered with structural biologists at Indiana University to conduct some experiments on Alda-1 to try to find out how it works.
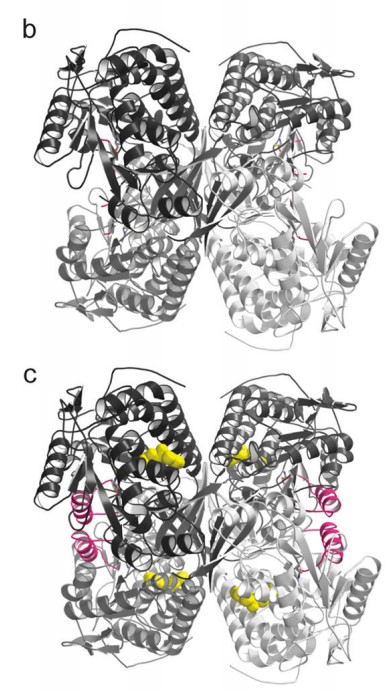
To do this, and why this paper was published in Nature Structural and Molecular Biology rather than another journal, they used a technique called X-ray Crystallography to “see” if (and how) Alda-1 interacts with ALDH2. Some of the results of these experiments are shown above. On the left, Panel B (on top) shows a 3D structure of the “defective’ version of ALDH2. If you’re new to structural biology papers, this will take some time getting used to it, but if you look carefully, you can see that ALDH2 is a tetramer: there are 4 identical pieces (in the top-left, top-right, bottom-left, bottom-right) which are attached together in the middle.
It’s not clear from this picture, but the defective version of the enzyme differs from the normal because it is unable to maintain the 3D structure needed to link up with a coenzyme (a chemical needed by enzymes which do this sort of chemical reaction to be able to work properly) called NAD+ or even carry out the reaction (the “active site”, or the part of the enzyme which actually carries out the reaction, is “disrupted” in the mutant).
So what does Alda-1 do, then? In the bottom (Panel C), you can see where the Alda-1 molecules (colored in yellow) are when they interact with ALDH2. While the yellow molecules have a number of impacts on ALHD2’s 3D structure, the most obvious changes are highlighted in pink (those have no clear counterpart in Panel B). This is the secret of Alda-1: it actually changes the shape of ALDH2, (partially) restoring the enzyme’s ability to bind with NAD+ and carry out the chemical reactions needed to process acetaldehyde, and all without actually directly touching the active site (this is something which you can’t see in the panel I shared above, but you can make out from other X-ray crystallography models in the paper).
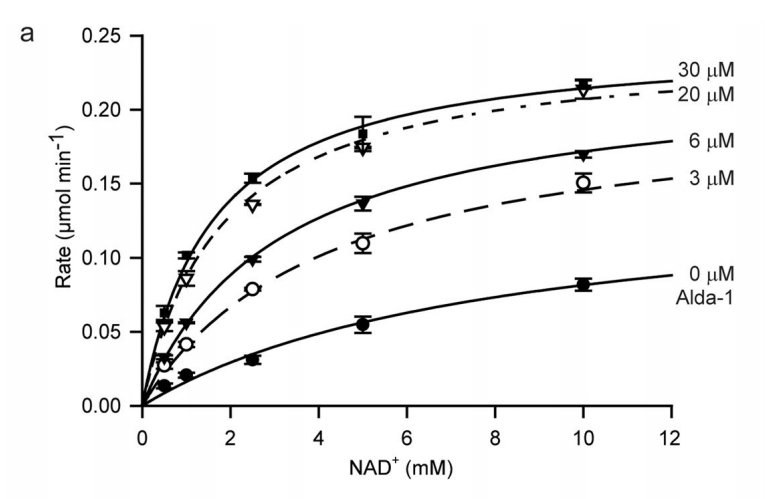
The result? If you look at the chart below (Panel A), you’ll see two relationships at play. First, the greater the amount of co-enzyme NAD+ (on the horizontal axis), the faster the reaction speed (on the vertical axis). But, if you increase the amount of Alda-1 from 0 uM (the bottom-most curve) to 30 uM (the highest-most curve), you see a dramatic increase in the enzyme’s reaction speed, for the same amount of NAD+. So, does Alda-1 activate ALDH2? Judging from this chart, it definitely does.
Alda-1 is particularly interesting because most of the chemicals/drugs which we are able to develop work by breaking, de-activating, or inhibiting something. Have a cold? Break the chemical pathways which lead to runny noses. Suffering from depression? De-activate the process which cleans up serotonin (“happiness” chemicals in the brain) quickly. After all, its much easier to break something than it is to fix/create something. But, instead, Alda-1 is actually an activator (rather than a de-activator), which the authors of the study leave as a tantalizing opportunity for medical science:
This work suggests that it may be possible to rationally design similar molecular chaperones for other mutant enzymes by exploiting the binding of compounds to sites adjacent to the structurally disrupted regions, thus avoiding the possibility of enzymatic inhibition entirely independent of the conditions in which the enzyme operates.
If only it were that easy (it’s not)…
Where should we go from here? Frankly, while the paper tackled a very interesting topic in a pretty rigorous fashion, I felt that a lot of the conclusions being drawn were not clear from the presented experimental results (which is why this post is a bit on the vague side on some of those details).
I certainly understand the difficulty when the study is on phenomena which is molecular in nature (does the enzyme work? are the amino acids in the right location?). But, I personally felt a significant part of the paper was more conjecture than evidence, and while I’m sure the folks making the hypotheses are very experienced, I would like to see more experimental data to back up their theories. A well-designed set of site-directed mutagenesis (mutating specific parts of ALDH2 in the lab to play around with they hypotheses that the group put out) and well-tailored experiments and rounds of X-ray crystallography could help shed a little more light on their fascinating idea.
Paper: Perez-Miller et al. “Alda-1 is an agonist and chemical chaperone for the common human aldehyde dehydrogenase 2 variant.” Nature Structural and Molecular Biology 17:2 (Feb 2010) –doi:10.1038/nsmb.1737
